Home Page for Ian Wilson
email: I.J.Wilson@ncl.ac.uk
R packages
Packages (with varying degree of completeness) for the R software system. Latest versions
in my R package directory
Install packages on unix/linux using the commands
R CMD INSTALL
packagename.tar.gz
To install on a windows systems use the zip versions
and install from within R using
> install.packages("packagename.zip",repos=NULL)
Where you have downloaded the package to your working directory. Once installed the
zip file can be safely deleted.
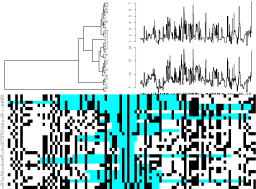
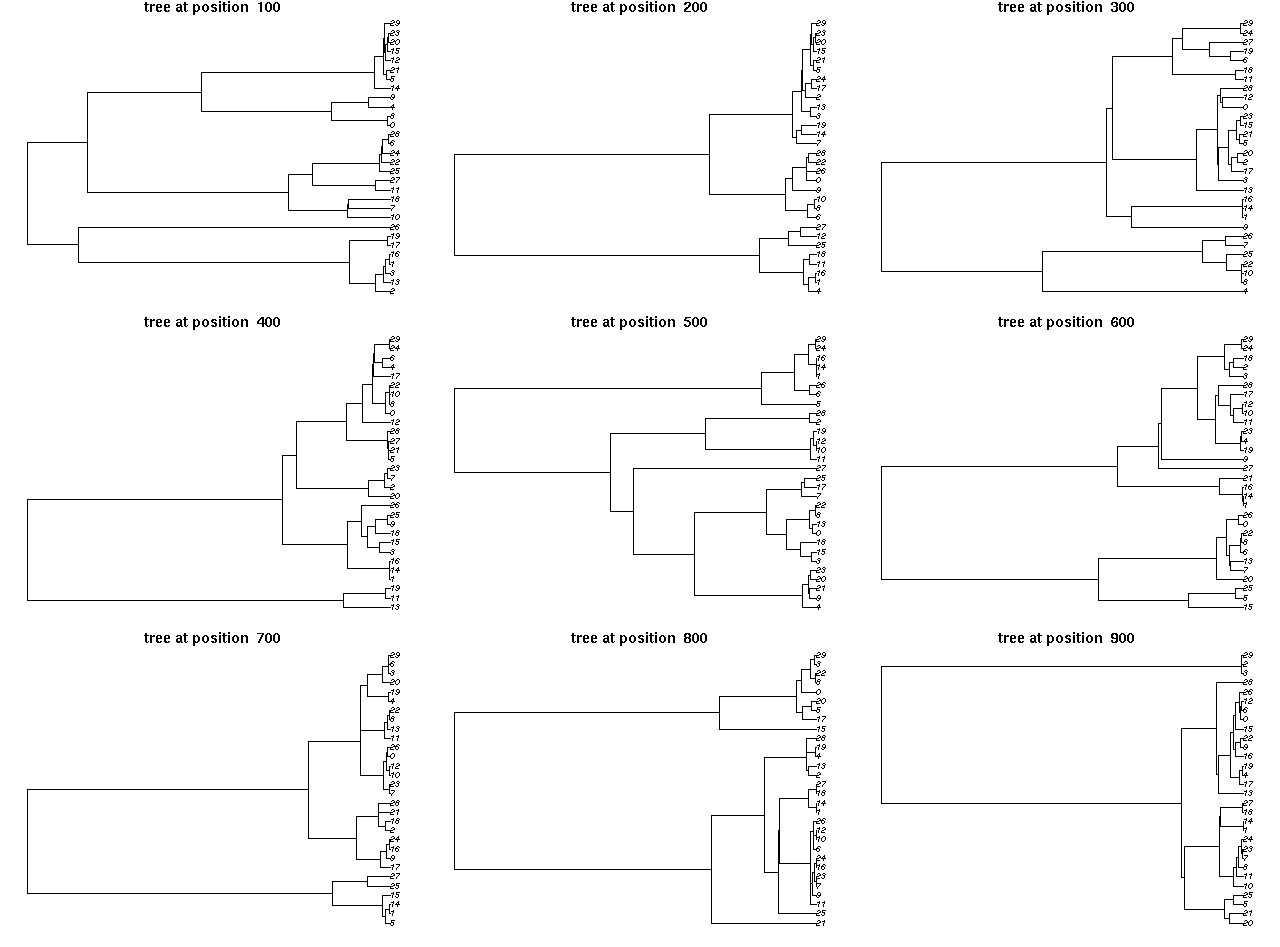
Screenshots of interactive plots from the ARG package. Left plot
shows a view of an ancestral graph showing genetic variants (bottom
panel) along with the tree under a single site (top left panel) and
tree heights and lengths (top right panel). Right plot gives views
of trees 100 sites apart. Click on either plot for a larger
view.
C/C++ Software
Unix Systems
All Software on this page is in the form of zipped tar files.
Use
gunzip -c filename.tar.gz | tar
xf -
then type make
to compile the program. A limited manual for micsat is in the file
micsat.doc. Other manuals
can be downloaded in pdf format.
Windows (95/98/NT)
The programs are distributed as executable files in a zip file
format
Macintosh (MacOS 8.0 or above)
A binhex-ed executable for the Macintosh, MacBatwing, may be
downloaded.
On launching, the program opens a dialog box with a text field. The
user puts the program arguments (infile outfile seed) into this
field and hits ok.
(Compilation on the Mac courtesy of Oliver Pybus - send
comments/bugs to I.Wilson@maths.abdn.ac.uk).
Program Descriptions
Micsat
 |
This is the program described in the paper "Genealogical
inference from microsatellite data" by Wilson and Balding published
in Genetics 150: 499-510.
A description of the program and
the downloaded files.
|
|
Bayesian Analysis of Trees With Internal Node Generation. Batwing is described in
Wilson, Weale & Balding 2003.Inferences from DNA data:
population histories, evolutionary processes and forensic match
probabilities. Journal of the Royal Statistical Society: Series
A, 166: 155-188.
Download |
 |
Parentage
Program described in Emery, Wilson, Craig, Boyle and Noble
(2001). Assignment of paternity groups without access to parental
genotypes: multiple mating and development plasticity in
squid.
Molecular Ecology: 10: 1265-1278.
download |
Hapgen
A program for haplotype reconstruction from genotypes. Work done with David
Cooper.
This program is a Markov chain Monte Carlo method to reconstruct
haplotypes from genotypes. It uses Metropolis coupled chains to
speed up mixing.
Includes options to deal with:
- Known phase,
- Missing Data,
- Multiple subpopulations
Ancestors
Inference of migration or population splitting using importance
sampling.
Vickers, McLeod, Spector & Wilson (2001). Assessment
of mechanism of skewed X-inactivation by analysis of twins.
BLOOD: 97:
1274-1281
 analysis of Twins
Multivariate Normal Model, and Twins Stem Cell Number
analysis of Twins
Multivariate Normal Model, and Twins Stem Cell Number
I.J.Wilson@ncl.ac.uk
Last Modified 28 February 2006





